
|
November 7, 2007 |
Drosophila 12 Genomes Consortium
Biologists Assemble Fly mtDNA for Landmark Genome Project
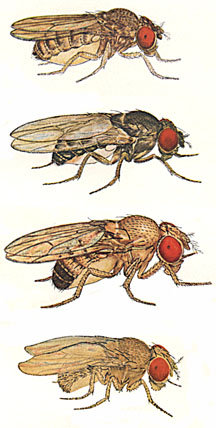
As part of a major new international genome sequencing project, Brown biologists assembled the complete mitochondrial DNA sequences of seven different species of fruit fly. Their work, published in Nature, provides scientists with an exciting new tool to understand the genetic differences within a species as well as the evolutionary relationships among different species. | |||
|
Brown University Home |
The work, appearing in Nature, is part of an international research effort to catalogue the DNA sequences of 12 species of Drosophila, or fruit fly, a critical and common laboratory model used to study human development, genetics, and evolution. Results of the “Drosophila Dozen” project are the complete sequences of both nuclear and mitochondrial DNA from the 12 species of fly – a data set that gives scientists the unprecedented ability to compare related species and how they changed over time. About 150 scientists from around the world, collectively known as the Drosophila 12 Genomes Consortium, came together to sequence, assemble and analyze the genomes, all from closely related species that range from Drosophila yakuba, a red-eyed variety found on the African savannah, to Drosophila mojavensis, a cactus-dweller from the Sonoran desert. David Rand, a professor in the Department of Ecology and Evolutionary Biology at Brown, along with postdoctoral research associate Kristi Montooth and laboratory technician Dawn Abt were the only scientists in the consortium to assemble mitochondrial DNA, or mtDNA, sequences. In the sequencing process, DNA is extracted, chopped into bits, then analyzed to determine the order of base pairs. These bits must then be put back together, or assembled, an exacting process that involves piecing together billions of base pairs in proper order using special computer software. The Brown team assembled mtDNA sequences from seven newly sequenced fruit fly genomes; MtDNA sequences from five other fly species were previously completed. Mitochondria are tiny machines inside cells that convert food energy into a form that can be used by the cell – making mitochondria the metabolic engines that feed every function of life, from breathing to thinking to moving. Although most DNA is found in the cell nucleus, mitochondria pack some DNA of their own. Much of this genetic material provides the instructions to make the proteins that combine oxygen with the energy stored in sugars to produce ATP, the cell’s main energy source. Rand, a member of Brown’s Center for Computational Molecular Biology, says mtDNA is a remarkable survivor from ancient times. Its origins are known to be bacterial and about 2 billion years ago it fused with the DNA from a different lineage of bacteria to create the ancestor of all modern cells. Since then, mtDNA has evolved in parallel with nuclear DNA – and much of it has been transferred to the nuclear genome. But 37 essential genes are retained in modern mitochondria. Because of the energy-producing reactions it carries out, mitochondria mutate their DNA at a rapid rate and are less able to “proofread” for these errors than genes in nuclear DNA. “That high mutation rate makes mtDNA a perfect tool for spotting genetic differences between individuals within a species and also for comparing closely related species,” Rand said. “As species become more distantly related, the number of differences in the mtDNA sequences gets larger. So mtDNA is a powerful, precise way to track ancestry.” By comparing the mtDNA sequences of the 12 fruit fly species, Rand and his team discovered that some genes, such as the COX family, are highly conserved, or change little over time. Other mtDNA genes, such as NADH, are fast-change artists and evolve rapidly. “The 12 Drosophila genomes give us an unprecedented opportunity to understand evolutionary adaptation right down at the genetic level,” Montooth said. “If we want to understand how the fly that lives on the savannah is different from the fly that lives in the desert, we can trace physiological differences back to specific genes. “At the same time,” Montooth said, “mtDNA can teach us about metabolic performance and how it can be disrupted through mutation, giving insight into possible mechanisms for mitochondrial diseases and conditions such as diabetes, deafness, and nerve damage that can result in vision loss or dementia.” The National Institute of General Medical Sciences funded the work. Editors: Brown University has a fiber link television studio available for domestic and international live and taped interviews and maintains an ISDN line for radio interviews. For more information, call the Office of Media Relations at (401) 863-2476. ###### | |||
 PROVIDENCE, R.I. [Brown University] — Brown
University biologist David Rand and members of his lab have made a major
contribution to a groundbreaking genome sequencing project –
single-handedly assembling the mitochondrial DNA sequences of seven species of
fruit fly.
PROVIDENCE, R.I. [Brown University] — Brown
University biologist David Rand and members of his lab have made a major
contribution to a groundbreaking genome sequencing project –
single-handedly assembling the mitochondrial DNA sequences of seven species of
fruit fly.