|
Undergraduate Study Computational Biology Undergraduate Program
Standard Program for the Sc.B. Degree Our program educates the student liberally in these fields, building on a foundation of coursework that may then focus via several possible tracks. The program offers four possible tracks: computational genomics, biological sciences, molecular modeling and applied mathematics and statistical genomics. The program requires a senior capstone experience that pairs students and faculty in creative research collaborations.
Computational Biology graduates are candidates for competitive positions in industry or for training in academic science.
Honors: To be a candidate for honors, a student must have a course record judged to be excellent by the concentration advisor and must complete a thesis judged to be outstanding by the faculty member supervising the work. At least four courses comprising a coherent theme in one of the following areas: 1. Biochemistry 2. Ecology 3. Evolution/Genetics 4. Neurobiology Two course from the following: CSCI 1950-L (Algorithmic Foundations of Computational Biology) PHP2620 (Statistical Methods in Bioinformatics) APMA 1660 (Statistical Inference II) BIOL 1430 (Computational Theory of Molecular Evolution)
The 10th Anniversary of
|
|||||
 |
Franco Preparata, An Wang Professor In the relatively remote past, the science of life concerned itself with macroscopic features of living matter, which were used to explain both the morphology and the functions of living organisms. |
|
Such features were the basis also for taxonomy of species, in that similarities, both morphological and functional, were construed as evidence of relatively close common ancestry on the evolutionary trail. Although the empirical laws of heredity, as well as the physical sites of the heredity carriers, had been known for a long time, the detailed mechanisms of the process remained nearly inscrutable until well into this century, except for the realization that certain chemicals, the nucleic acids, were intimately connected with the process. Scientific discoveries of about half a century ago were bound drastically to subvert this established mode of scientific inquiry. Crucial was the elucidation of the structure of DNA (deoxyribonucleic acid) and of its role as the fundamental carrier of hereditary information. The revolutionary discovery of the DNA doublehelix structure by Watson and Crick in 1953 ushered in the era of molecular biology. They showed that DNA is a sequence of pairs of four structurally similar basic constituents called bases and denoted by the standard letters A, C, G, and T (as is well known, these are the initials of their respective chemical denominations). In fact, each base can be paired (has strong chemical affinity) with just another base, so only the pairs AT, CG, GC, and TA occur in the DNA sequence. While this view of DNA is perfectly adequate for its description, it is perhaps more significant to consider DNA as the pairing of two complementary strands, each carrying the same hereditary (genetic) information. This structure is essential for DNA replication, the archetypal phenomenon of reproduction: the two strands of a sequence are separated in the cell and each of them is copied into a complementary strand, giving rise to two replicas of the original sequence. This brief and very schematic digression is not intended to oversimplify marvelously complex biological phenomena, but simply to provide a glimpse into the emerging discrete structure of molecular biology. In fact, the realization that the above four bases are the building blocks of the description of the genetic patrimony is appropriately viewed as the informatization of biology, in that it shifts the description into the conventional computer-science nomenclature of sequences over a finite alphabet. This feature is not exclusive to DNA, but recurs for other biomolecules, such as RNA (the other fundamental nucleic acid) and, with a larger alphabet, for proteins. This characterization establishes a natural link between the two domains, since they use analogous descriptive devices. Contemporary with the beginning of molecular biology was the advent of the computer era. In its first decade, the rather rudimentary technology made the computer seem more a wondrous curiosity than a tool accessible to vast segments of users. The physical size of the installations, the associated physical plant, and the dismally poor reliability of the computers in no way let anyone suspect its ubiquitousness today. Thus it is not surprising that contacts between biology and computer science materialized somewhat later.
As the computer field was progressing rapidly (already in the sixties was computer
science identified as an autonomous academic discipline), the informatization of
biology revealed an entirely new host of problems. The notions of morphological
or functional similarity evolved into the notion of similarity between sequences
(polymers) of chemical constituents. For any class of homologous such sequences
(we mean here just that two sequences are homologous if they can be meaningfully
compared) this approach immediately poses two problems. The first is the definition
of the metrics, i.e., a quantitative model for the measurement of sequence similarity
(or distance). The second is the development of methods (algorithms) to carry out
such quantitative assessment.
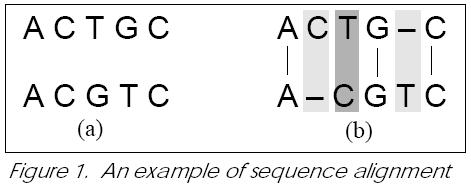
This trivial example illustrates the features of the alignment problem, which plays
an important role in computational biology, both for its modeling difficulties (which
are the biologically significant primitives and their weights?) and its algorithmic
complexity (the collective alignment of several sequences of several hundred characters
each). In addition to alignments, a vast collection of problems lies today at the
intersection of computer science and biology: DNA fragment assembly, physical mapping,
phylogeny, molecular structure prediction, genome rearrangement, and so on. Many
of these problems have been stimulated by the human genome project (i.e., the mapping
of the entire human DNA patrimony) but have also been fueled by rapidly growing
industrial interest.
Typically, computational biologists have been professionals from either field who
have taken on the difficult task of retraining themselves in the other discipline.
This approach has several shortcomings. First, individuals willing to undertake
such an unconventional educational path (self-teaching) are not the norm, and thus
are not numerous enough to fill a clearly identified professional need. Second,
a selfinstruction plan may not be sufficiently systematic to meet the requirements.
Third, one must take into consideration a subtle feature in the sociology of peer
groups in research and professions, that is, "acceptance". A peer group accepts
individuals with similar academic backgrounds, and such acceptance is rarely complete
in the case of retraining. Indeed, it has been quite common for computer science
to force (unrealistic) modelings for the benefit of algorithmic simplicity, and
for biologists to content themselves with commercially available software tools,
rather than undertaking original algorithmic development. This state of affairs
and the propitious opportunity offered by the excellent flexibility of Brown's undergraduate
curriculum prompted the idea of proposing a new undergraduate concentration in computational
biology. The core offerings of the concentration are designed to provide a balanced background in the interacting disciplines. This core is complemented by specialized tracks designed to differentiate among a number of related professions with identifiable expertise and skill. Thus the software track is for students interested in developing commercial software for biological applications; the molecular modeling track is for students interested in competence in molecular modeling and drug design; the biological sciences track is for students interested primarily in biological questions. In addition to core courses and electives, the program requires as its capstone experience the completion of a senior research project under close faculty supervision. The stewardship of the program is entrusted to three concentration advisors, respectively in Computer Science, Chemistry, and Biology and Medicine. Beside the usual advising responsibilities, the advisors have the task of supervising the evolution of the program.
Minimally, the proposed program identifies among the current offerings an instructional
package that can be legitimately named Computational Biology. As the program evolves,
we expect the establishment of a permanent lecture series and of graduate research
and instructional initiatives, and, possibly, the addition of faculty clearly identifiable
with the field. |
|
_______________________________________________________ top of page