Face-centered cubic (FCC) lattice models for protein folding
Welcome to Allan Stewart's protein folding thesis. The thesis was completed April 2010 for undergraduate honors in Computational Biology (Sc. B. program, Brown University).
download the thesis
download the presentation (ppt)
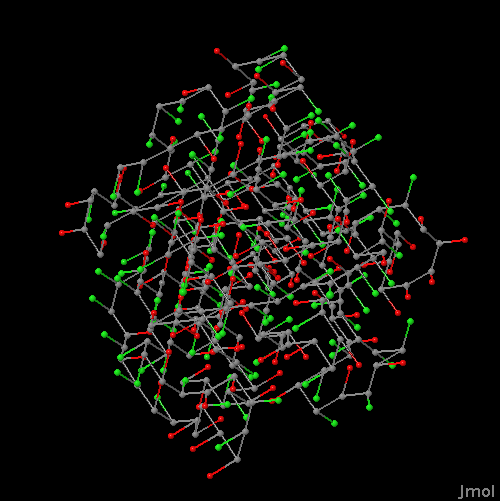
To view the dataset of 1198 PDB proteins on an FCC sidechain model with red hydrophobic sidechains and green hydrophilic sidechains, click here:
FCC PDB protein set
To view the Mathematica notebook containing a near-octahedron conformation on FCC, click here: FCC Mathematica 6 notebook (view in browser)
To view the pairs of amino acids most enriched in contacts, click here:
Most contact residue pairs
For text files listing PDB IDs found to be biplanar or non-biplanar, click here:
Biplanar ids, Non-biplanar ids
Thanks and Additional Software
Thanks very much to the Informatics group at the University of Freiburg for writing the LatFit software, which fits PDBs to the FCC sidechain lattice.
Additional software which may be of use:
- Protein visualization software by Chris Maloney (former Istrail Lab member) which interactively lets users place residues on 2D square, 3D cubic and FCC Lattice
- Ramachandran aligner software which finds Ramachandran angles which are energetically favorable for related protein species. This is a collaboration with Sorin Istrail and Yash Thakore.