- Genomic Regulatory Networks and the Regulatory Genome
- Computational Models of SNPs and Disease Associations
- CELLARIUM: An Integrated Framework for Genome Browsers and Bioinformatics Workflows
- Algorithmic Theory of Protein Folding and Misfolding
- Statistical Mechanics and Computational Complexity
- Education and the Classroom
This work was funded by the National Science Foundation

Preterm labor facts
- Occurs once in every 8 women.
- Estimates from twin studies suggest up to 40% heritability
- Genetic estimates put all of the risk on the maternal side
- More common in African Americans
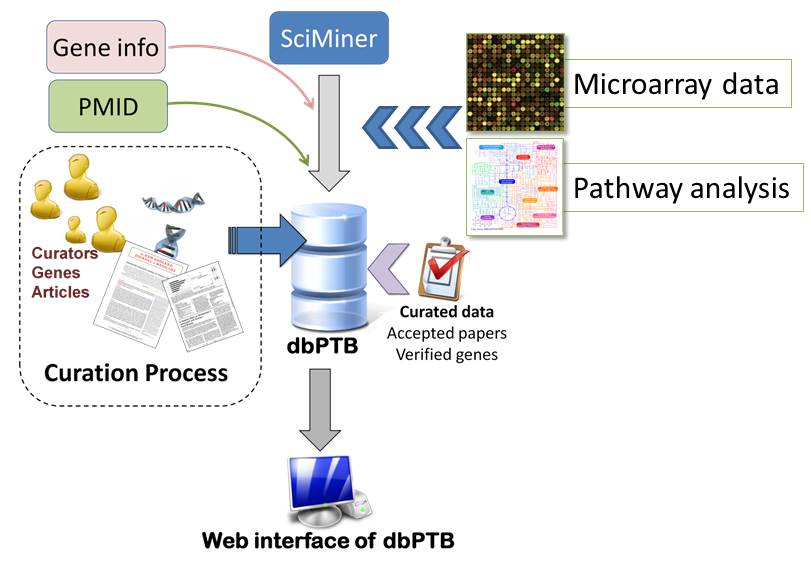
We are interested in the genetic architecture of preterm birth and have developed an approach to identify a more manageable set of candidate genes which nonetheless incorporates some elements of genome wide investigation for the study of preterm birth. Our approach combines information from published literature with data from expression databases, linkage data and pathway analyses to identify biologically relevant genes for testing in an association study of genetic variants and preterm birth. We imported data from publicly available array databases like expression arrays from the placenta of patients that delivered preterm versus those who delivered at term and similar transcription wide expression data from uterine tissue from preterm and term delivery. The last step was pathways analysis. We included genes that were strongly represented in the pathways even if not identified by the microarrays or literature search.
For more information contact Alper Uzun
Publications
| 2013 | |
| [2] | Alper Uzun, Andrew T. Dewan, Sorin Istrail, James F. Padbury, "Pathway-based genetic analysis of preterm birth", In Genomics, vol. 101, no. 3, pp. 163 - 170, 2013. [bib] [pdf] [doi] |
| 2012 | |
| [1] | Alper Uzun, Alyse Laliberte, Jeremy Parker, Caroline Andrew, Emily Winterrowd, Surendra Sharma, Sorin Istrail, James F. Padbury, "dbPTB: a database for preterm birth", In Database (Oxford), vol. 2012, pp. bar069, 2012. [bib] [pdf] [doi] |
