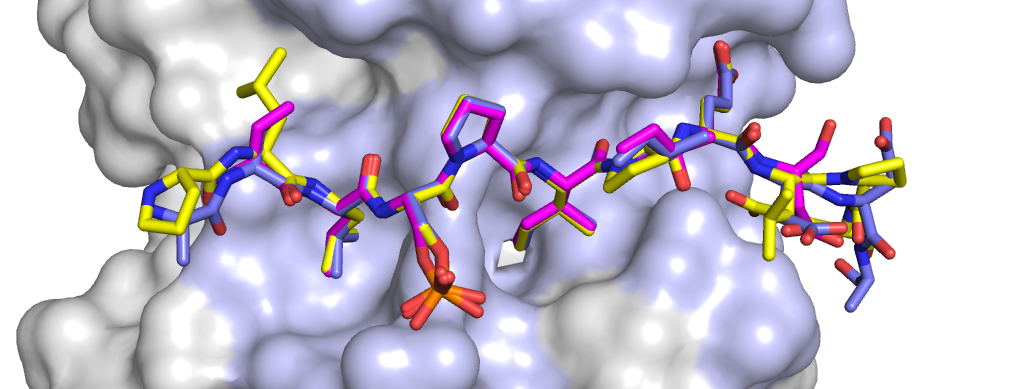
Xinru Wang, et. al./Brown University
PROVIDENCE, R.I. [Brown University] — Proteins often don’t do anything in our cells by themselves. Their role in a given physiological process – good or bad – is typically determined by binding to one or more other proteins. So as researchers try to figure out why the enzyme PP2A is implicated in many cancers, a new study provides a potentially significant break in the case. It identifies 98 proteins that are very likely to bind with PP2A, making them potential partners in crime worth investigating.
The discovery is the result of a detailed study of PP2A’s structure down to the atomic level by a team of four scientists at Brown University and one in Leuven, Belgium. By determining exactly how PP2A physically binds with other proteins, the scientists were able to figure out which other proteins are a strong fit. Most of the proteins on the list are news to the biological community.
Moreover, the study data could help scientists to predict how strongly each protein, or “substrate” binds PP2A. It could also offer clues on how to prevent a binding if it turns out to be disease-causing, for instance because it misregulates a process in a cell.
“PP2A is a very important enzyme, but identifying its substrates and its regulators has been exceptionally difficult and part of the reason is we didn’t understand how they were engaging with this protein,” said corresponding author Rebecca Page, a Professor of molecular biology, cellular biology and biochemistry at Brown University and corresponding author of the paper in the journal Structure.
Binding findings
Brown graduate student Xinru Wang and postdoctoral researcher Rakhi Bajaj led the effort to find PP2A’s closest binding partners. Their main strategy was to bind the regulatory subunit of PP2A, called B56, with three shortened versions of two proteins already known to be associated: “BubR1” and “RepoMan.” They made each of those protein complexes into crystals, which can be analyzed in atomic detail using X-rays. This procedure gave them a close look at which key binding interactions were consistent and which other interactions made each union unique.
“The similarities were the common features that we used to identify additional PP2A interactors,” Wang said.
And, Page added, “The differences between RepoMan and BubR1 was what allowed us to explain why the binding affinities were different and how different PP2A regulators might be competing in the cell.”
One of the key similarities they observed, for example, was that PP2A has two distinct pockets at a key location on its surface where other proteins can dock if they have the right protrusions.
A key difference they noticed is that RepoMan’s binding is much stronger than that of BubR1 and they could discern structurally why that was – RepoMan builds an extra “salt bridge” to solidify its connection.
Making a list
The researchers used these observations and others made from studying the three crystallized bindings to guide a search of databases of human proteins. By looking for specific amino acid sequences in proteins the scientists were able to tell how well they might fold up and bind to PP2A.
Page said that with conservative criteria – looking only for the best and most likely matches – they found 98 proteins with strong likelihoods. A few of them were already known to associate with PP2A – confirming that the approach makes sense —but many more are novel.