PROVIDENCE, R.I. [Brown University] — Scientists report in a new study that by imitating a natural process of cells, they prevented the formation of protein clumps associated with amyotrophic lateral sclerosis (ALS) and frontotemporal dementia.
In lab cultures of human and yeast cells, the scientists stopped the harmful clumping of FUS proteins by exposing them to phosphorylation, a process that makes precise changes to the amino acid building blocks of proteins, increasing their negative electric charge. The research shows that the increase in charge causes the proteins to repel when they normally might aggregate.
The findings could eventually have positive implications for the treatment of ALS — commonly called Lou Gehrig’s disease — and dementia.
“No one has shown that you can use charge, and phosphorylation as a way to get charge, to disrupt these ALS-associated protein aggregates,” said co-corresponding author Nicolas Fawzi, an assistant professor in the Department of Molecular Pharmacology, Physiology and Biotechnology at Brown University.
Phosphorylation occurs in cells throughout the body for various reasons. Cells perform it on FUS proteins as part of a sequence of events related to DNA damage. In the new study, the researchers examined both natural phosphorylation and artificial phosphorylation by replacing specific amino acids in FUS proteins.
The study, conducted by a team of researchers at Brown, the Uniformed Services University, Johns Hopkins University, Lehigh University and the National Institutes of Health, appears in the EMBO Journal.
Study steps
The project began in Fawzi’s lab at Brown, where he’s been studying the structure and biophysics of FUS and similar proteins closely associated with neurodegenerative diseases. He and co-lead author Veronica Ryan, a Brown neuroscience graduate student, wanted to explore how and whether phosphorylation would affect the way FUS proteins can clump together, which undermines the health of neurons.
The first task was to identify several specific amino acid sites in the region FUS uses to bind with other proteins where the natural enzyme DNA-dependent protein kinase (DNA-PK) causes phosphorylation.
Their collaborators at the Uniformed Services University, including co-lead author Zachary Monahan and co-corresponding author Frank Shewmaker, continued the work by confirming these and more phosphorylation sites in FUS derived from human cells.
With more than a dozen sites identified, the team then set out to imitate the work of DNA-PK in living cell cultures. Monahan and Shewmaker’s team engineered human FUS into brewer’s yeast and also implemented the phosphorylation mimicry in a culture of human cells.
What they observed in both cultures is that by increasing the degree of phosphorylation — from just two sites all the way up to 12 — they could proportionally reduce the propensity of the protein to aggregate into clumps.
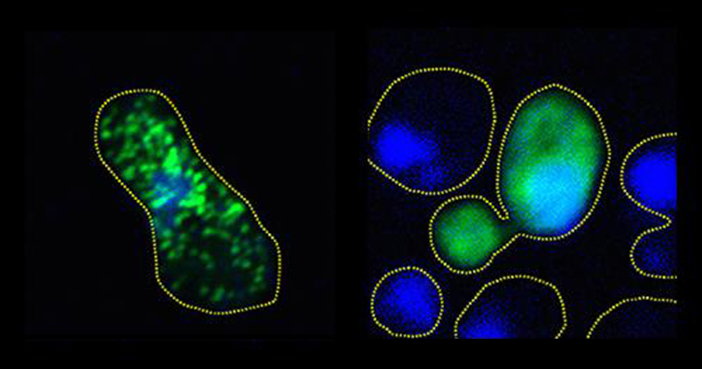
In the yeast cells, they showed that the more effectively they prevented full aggregation — by adding more sites mimicking phosphorylation — the more robustly colonies of the cells would grow. That finding demonstrated that FUS clumping is toxic to cells.
Ryan, Abigail Janke and colleagues at Brown showed how phosphorylation disrupts contacts between molecules of FUS. Meanwhile, computational modeling of the effect of the phosphorylation in the structure of the protein, led by co-author Jeetain Mittal at Lehigh, showed that the change in electric charge is what led to the difference in the proteins’ likelihood of aggregating.
A therapeutic horizon
Fawzi acknowledged that the approach the scientists used in the lab would not yet constitute a practical therapy for neurodegenerative diseases, but he said the demonstration that phosphorylation can be used to disrupt protein aggregation should motivate work to produce a treatment.
“There’s no therapy or cure for ALS and frontotemporal dementia,” Fawzi said. “What we need are new hypotheses and new angles.”
Fawzi hypothesized that because phosphorylation is a natural process of cells, the key may be finding a “switch” to turn it on when needed. He also noted that many pharmaceutical companies have very active research groups dedicated to harnessing the phosphorylating enzymes known as kinases like DNA-PK.
A future therapy, he said, would have to be targeted — localized in the brain in the case of frontotemporal dementia, or focused on motor neurons in the case of ALS. Phosphorylating FUS in the body more generally could be harmful, given that it is presumed to have important roles in helping cells process RNA and may have a role in repairing DNA.
For now, the collaboration is focused on testing phosphorylation specifically in a model of a neurodegenerative disease, rather than in cells in general, Fawzi said.
The paper’s other authors are Kathleen Burke, Shannon Rhoads, Guz Zerze, Robert O’Meally, Gregory Dignon, Alexander Cnicella, Wenwei Zheng, Robert Best and Robert Cole.
The National Institutes of Health (R01GM118530, R35GM119790, P20GM104937), the DEARS Foundation and the Rhode Island Foundation funded the research.