PROVIDENCE, R.I. [Brown University] — The field of mathematical topology is often described in terms of donuts and pretzels.
To most of us, the two differ in the way they taste or in their compatibility with morning coffee. But to a topologist, the only difference between the two is that one has a single hole and the other has three. There’s no way to stretch or contort a donut to make it look like a pretzel — at least not without ripping it or pasting different parts together, both of which are verboten in topology. The different number of holes make two shapes that are fundamentally, inexorably different.
In recent years, researchers have drawn on mathematical topology to help explain a range of phenomena like phase transitions in matter, aspects of Earth’s climate and even how zebrafish form their iconic stripes. Now, a Brown University research team is working to use topology in yet another realm: training computers to classify how human cells organize into tissue-like architectures.
In a study published in the journal Soft Matter, the researchers demonstrate a machine learning technique that measures the topological traits of cell clusters. They showed that the system can accurately categorize cell clusters and infer the motility and adhesion of the cells that comprise them.
“You can think of this as topology-informed machine learning,” said Dhananjay Bhaskar, a recent Ph.D. graduate who led the work. “The hope is that this can help us to avoid some of the pitfalls that affect the accuracy of machine learning algorithms.”
Bhaskar developed the algorithm with Ian Y. Wong, an assistant professor in Brown’s School of Engineering, and William Zhang, a Brown undergraduate.
There’s been a significant amount of work in recent years to use artificial intelligence as a means of analyzing big data with spatial information, such as medical imaging of patient tissues. Progress has been made in training these systems to classify accurately, “but how they work is opaque and a little finicky,” Wong said. “Just like people, sometimes computers hallucinate. You can have a few pixels in the wrong place, and it can confuse the algorithm. So Dhananjay has been thinking about ways we might be able to make those analyses a little more robust.”
In developing this new system, Bhaskar took inspiration from modern art, specifically Pablo Picasso’s “Bull.” The series of 11 lithographs starts with a bull depicted in full detail. Each successive frame strips away a bit of detail, ending in a simple drawing capturing only the animal’s fundamental attributes. By employing topology, Bhaskar thought he might be able to do something similar to understand the underlying form of tissue-like architectures.
The way in which cells migrate and interact depends on the physiology of the cells involved. For example, healthy tissues contain higher numbers of stationary epithelial cells. Processes like wound repair or cancer, however, often involve more mobile mesenchymal cells. Differences in physiology between the two cell types cause them to cluster together differently. Epithelial cells tend to aggregate into larger, more closely packed clusters. Mesenchymal cells tend to be more dispersed, with groups of cells branching off in different directions. But when assemblages contain a mix of both kinds of cells, it can be difficult to accurately analyze them.
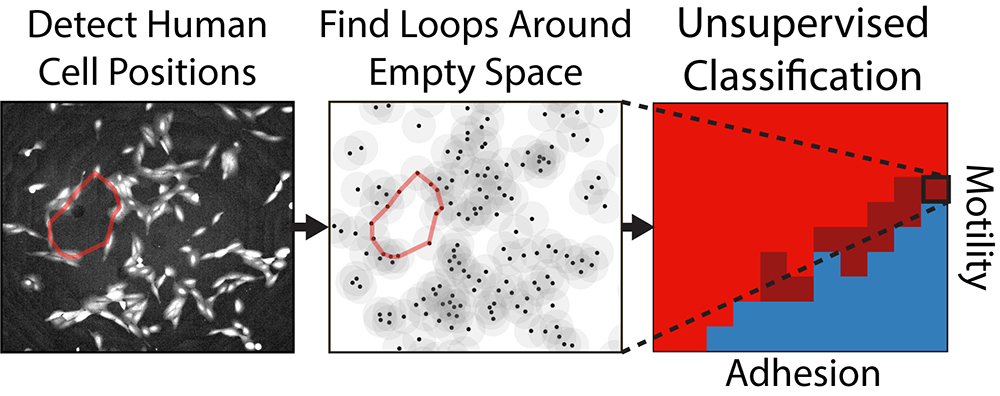
The new algorithm uses a mathematical framework called persistent homology to examine microscope images of cell assemblages. Specifically, it looks at the topological patterns — loops or holes — that the cells form collectively. By looking at which patterns persist across different spatial resolutions, the algorithm determines which patterns are intrinsic to the image.
It starts by looking at the cells in their finest detail, determining which cells seem to be part of topological loops. Then it blurs the detail a bit by drawing a circle around each cell — effectively making each cell a little larger — to see which loops persist at that more coarse-grained scale and which get blurred out. The process is repeated until all the topological features eventually disappear. At the end, the algorithm produces a sort of bar code showing which loops persist across spatial scales. Those that are most persistent are stored as a simplified representation of the overall shape.
As it turns out, those persistent topological objects can be used to categorize clusters of differing types of cells. After training their algorithm on computer-simulated cells programmed to behave like different types of cells, the team turned it loose on real experimental images of migratory cells. Those cells had been exposed to varying biochemical treatments so that some were more epithelial, some were more mesenchymal, and some were somewhere in between. The study showed that the topological algorithm was able to correctly classify different spatial patterns according to which biochemical treatment the cells had received.
“It was able to pull out all of these experimental treatments just by identifying these persistent topological loops,” Wong said. “We were kind of amazed at how well it did.”
The team hopes that one day the algorithm could be used in laboratory experiments to test drugs, helping to determine how different drugs can alter cell migration and adhesion. Eventually, it may also be used on medical images of tumors, potentially helping doctors to determine how malignant those tumors may be.
“We’re looking for ways to catch subtleties that might not be apparent to the human eye,” Wong said. “We hope that this might be a human interpretable approach that complements existing machine learning approaches.”
The research was supported by National Cancer Institute’s Innovative Molecular Analysis Technologies Program (R21CA212932) and the Brown University Data Science Initiative.