Complex organisms are built from simple starting blocks, and, for Brown University neuroscientist Kate O’Connor-Giles, the brain is the most fascinating piece in this puzzle.
“I’m still awed by the fact that it’s possible to generate a nervous system that’s capable of controlling all the complex behaviors we see around us and gives rise to human thought and emotion through concrete processes that can be understood by scientific investigation,” said O’Connor-Giles, provost's associate professor of brain science and associate professor of neuroscience.
O’Connor-Giles, who is particularly interested in understanding how neurons communicate, says her research has already moved in new directions through interdisciplinary collaborations since joining Brown’s Robert J. & Nancy D. Carney Institute for Brain Science in July 2018.
She recently took a break from designing and carrying out scientific experiments to chat about basic research and new CRISPR-based gene editing tools developed by her lab to advance the study of synapses in vivo.
Q: How did you become interested in studying how neurons communicate and give rise to thought and behavior?
A: As a graduate student first in Chris Doe’s lab, then in Jim Skeath’s lab at Washington University in St. Louis, my work focused on understanding how the great diversity of neurons and glia in the brain is generated during development. I loved this area of research as it directly addressed a particularly fascinating aspect of the question that had sparked my interest for so long: ‘how do complex organisms arise from simple building blocks?’.
As a postdoctoral researcher, I decided to continue studying neural development, but with a focus on later events in the building a functional nervous system. I joined Barry Ganetzky’s lab just as they had found a number of genes that regulate synaptic growth, and, building on these discoveries, investigated how signaling between pre- and post-synaptic cells coordinates synapse formation and communication between the two cells.

My lab’s focus has evolved further into working to understand how synapses are organized to achieve specific properties and how they can be changed after development to alter those properties. We believe that answering these questions will help uncover how the nervous system can ensure robust, predictable communication while remaining malleable enough to continuously incorporate new experiences. Importantly, it’s now apparent that many neurological disorders can be traced to aberrant synapse formation and function, so we are interested in understanding how these processes can go awry.
Q: Tell us about one project you’re currently working on, and why you're particularly excited about it.
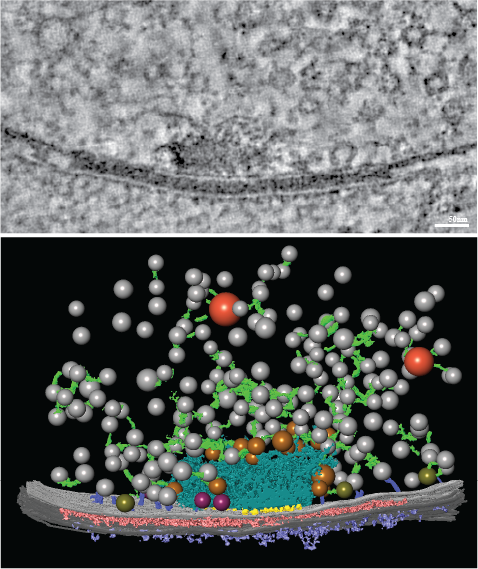
A: We have been investigating the regulation of calcium channels as synapses adapt to various perturbations. Synaptic communication involves the release of neurotransmitter by the presynaptic neuron that then binds neurotransmitter receptors in the postsynaptic neuron. Neurotransmitter release depends on calcium, so calcium influx through presynaptic calcium channels plays an important role in defining communication between neurons.
At Drosophila (fruit fly) motor synapses, we know through the work of others that calcium influx is modulated to alter neurotransmitter release as neurons adapt to insults, but we don’t know how. For example, when flies are exposed to a wasp toxin that rapidly impairs neurotransmitter receptors, neurotransmitter release is increased to precisely compensate for the toxin’s effect. By analogy to a conversation—when the postsynaptic, or ‘hearing,' side of the synapse is impaired, that cell sends a signal to the presynaptic neuron, triggering it to 'speak' louder.
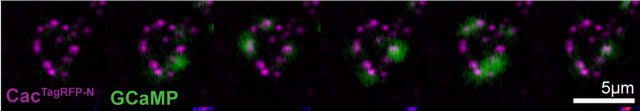
To understand how calcium influx is tuned during this and related processes, we used CRISPR to insert a fluorescent tag into the gene that encodes the fly calcium channel responsible for neurotransmitter release. We’re really excited about this because it enables us to visualize calcium channel dynamics during adaptation to perturbations for the first time.
In live imaging experiments, we found that more calcium channels accumulate at presynaptic terminals within 10 minutes of exposure to wasp toxin. Together, with work from other labs, this tells us that synapses can undergo global structural remodeling on very short timescales to enhance neurotransmitter release when needed. Interestingly, with our collaborators Dion Dickman and Pragya Goel, we find that when neurotransmitter release needs to be decreased to compensate for destabilizing changes in the opposite direction, calcium influx is modulated through a different mechanism. So, these neurons have a repertoire of strategies they tap into for tuning neurotransmitter release to maintain stable communication.
Q: Your lab has developed new genome engineering tools to overcome barriers to the study of synapses in organisms. What are some of these barriers, and how do the genome engineering tools help overcome them?
A: Drosophila is an ideal model system for genetic studies of the nervous system in many ways. It has powerful genetic tools; a manageable number of neurons and synapses that control complex behaviors; well-studied model synapses, circuits and behaviors; and synaptic genes that are conserved in humans. However, until recently it has been very challenging to make targeted edits to the fly genome. By targeted edits, I mean introducing new sequences—such as the fluorescent tag I mentioned earlier—into a specific site in the genome, precisely mutating one or more genes to probe gene function, or introducing disease-associated mutations for studying their effects.
We were working on ways to improve this when the first reports of CRISPR-based gene editing came out. We immediately formed a collaboration with Jill Wildonger and Melissa Harrison at the University of Wisconsin-Madison to adopt CRISPR in flies. This, and work from many other labs, has made gene editing remarkably straightforward in flies today.
We've since focused on developing approaches that make CRISPR-based screening feasible in vivo. We’re also using genome engineering for detailed studies of gene function and regulation. For example, in a collaborative project with Karla Kaun’s lab, we're using CRISPR to understand how alcohol affects gene expression. Many genes can be expressed in multiple, distinct forms, and work from the Kaun Lab recently showed that alcohol alters which form is used. Figuring out how these changes affect brain function requires altering the regulation of targeted genes in their normal genomic context, which would have been extremely difficult to accomplish without an efficient way to edit the Drosophila genome.
Q: How does your research provide insight into particular neurological disorders?
A: We’re conducting genetic screens to identify and characterize new synaptic genes that are conserved from flies to humans. A number of the genes we’ve identified are associated with genetic risk for neurological disorders, including mental health disorders and neurodegeneration. The fly is a powerful model system that we use to figure out how these genes normally act in the nervous system and how disrupting their function impairs synaptic communication. With respect to our project looking at the compensatory changes that neurons make to ensure normal communication is maintained in the face of perturbations, it is our hope that by figuring out exactly how neurons achieve this, we will identify targets for therapeutic interventions that could be used to modulate synaptic function during disease.